|
While contrast enhancement makes it easier to detect cFos-IR cells, it can result in an overestimation of total cell count due to increased pixel intensity of dimly-stained cells. Edge detection allows for clean segmentation of cells within colonies but can detect false edges in background staining present with cFos cells which can obscure the cells of interest. However, these algorithms encounter problems with edge detection, contrast enhancement and denoising in brain tissue analysis 14, 15, 16, 17, 18. Additionally, as the number of images increases so does fatigue and the potential for increased errors in counts.Įxisting automated methods can be used to quantify cFos-like cells even though they are optimized for cell colony or tumor spheroid analysis. While analysis with select software improves the reliability of manual quantification, an experimenter must still determine whether to count a dark-stained cell based on smoothness, clarity and size. Alternatively, digital images of cFos-stained tissue can be obtained, imported and thresholded using software such as ImageJ to aid in counting 4, 5, 12, 13. For manual quantification, cells can be counted in areas within a set microscopic field of view. Brain slices are typically immunohistochemically stained for cFos protein, resulting in round, light to dark-labeled cells. To analyze cFos-IR cells, experimenters can choose between manual or automated methods for quantification. 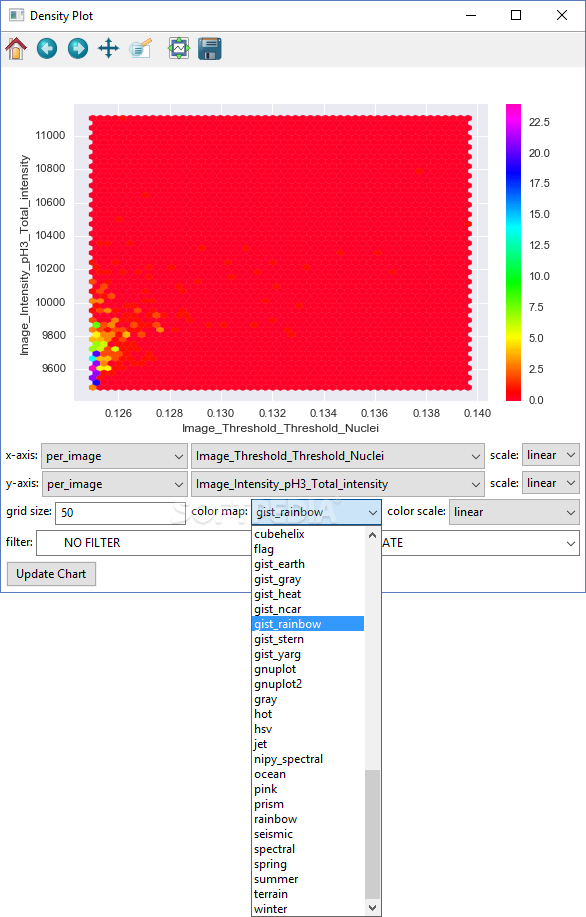
For example, characterization of the brain regions and cell types in which cFos protein is increased has provided insight into the cell populations that contribute to learning and memory, drug addiction, obesity and fear conditioning 3, 4, 5, 6, 7, 8, 9, 10, 11. Behaviorally-induced increases in the number of cFos-Immunoreactive (cFos-IR) cells often suggest that these activated neuronal ensembles may contribute to specific behaviors. cfos mRNA is transcribed within minutes of stimulus exposure and results in the cytoplasmic expression of cFos protein 60–90 min later 2. cFos, a commonly studied IEG, is a proto-oncogene and member of the Fos family of transcription factors. Immediate early genes (IEGs) are rapidly transcribed and translated upon stimulus exposure, making them useful for post-behavioral, correlated readouts of cellular activity 1. The current study demonstrates that SCC is a novel, automated tool to quantify cells in brain tissue and complements current, open-sourced methods designed to detect cells in vitro. Additionally, SCC utilized a new approach to count overlapping cells with a pretrained convolutional neural network classifier. We found SCC to be highly accurate and efficient in quantifying cells with circular morphologies that expressed cFos. 
In Experiment 3, SCC analysis was conducted on images it was not trained on, to assess its general utility. In Experiment 2, performance analytics of OCFU, IMJM and SCC were compared. The absolute error in counts (manual versus automated method) was calculated and error types were categorized as false positives or negatives. In Experiment 1, manually-obtained cell counts were compared to those detected via OCFU, IMJM and SCC. Therefore, we created SimpylCellCounter (SCC), an automated method to quantify cells that express cFos protein, an index of neuronal activity, in brain tissue and benchmarked it against two widely-used methods: OpenColonyFormingUnit (OCFU) and ImageJ Edge Detection Macro (IMJM). Manual quantification of activated cells can provide valuable information about stimuli-induced changes within brain regions however, this analysis remains time intensive.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
